|
|
I
|
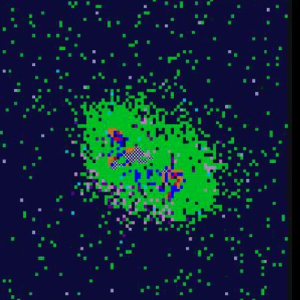
Identifying control mechanism of granuloma formation
during M. tuberculosis infection using an agent based model
|
|
|
Jose L. Segovia-Juarez, Suman Ganguli, and Denise Kirschner,
Journal of Theoretical Biology. Vol. 231, Issue 3, pp.357-376, 2004.
|
|
Granuloma Time-Lapse
Simulations
|
|
|
II
|
A comparison of random vs. chemotaxis-driven contacts of T-cells
with dendritic cells during repertoire scanning
|
|
|
Thomas Riggs, Adrienne Walts, Nicolas Perry, Laura Bickle,
Jennifer N. Lynch, Amy Myers, Joanne Flynn, Jennifer J. Linderman,
Mark J. Miller, and Denise E. Kirschner,
Journal of Theoretical Biology., Vol. 250, Issue 4, pp.732-751, 2008.,
doi:10.1016/j.jtbi.2007.10.015, PMID: 18068193, PMCID: PMC2548315 |
|
T cell Chemotaxis Time Lapse Simulations
|
|
|
III
|
Characterizing the dynamics of CD4+ T cell priming within a lymph node
|
|
|
Jennifer J. Linderman, Thomas Riggs, Manjusha Pande, Simeone Marino,
and Denise E. Kirschner, Journal of Immunology, 2010, 184: pp 2873-85,
doi:10.4049/jimmunol.0903117, PMID: 20154206, PMCID: 3153313
|
|
LN T-zone Time Lapse Simulations
|
|
|
IV
|
A Synergy between Individual TNF-Dependent Functions Determines Granuloma
Performance for Controlling Mycobacterium tuberculosis Infection
|
|
|
J. Christian Ray, Joanne L. Flynn, and Denise Kirschner, Journal of Immunology, 2009, 182: pp 3706-3717,
PMID: 19265149, PMCID: 3182770
|
|
Granuloma Time-Lapse
Simulations
|
|
|
V
|
Multiscale computational modeling reveals a critical role for TNF-α receptor 1
dynamics in tuberculosis granuloma formation
|
|
|
Fallahi-Sichani M, El-Kebir M, Marino S, Kirschner DE, Linderman JJ.,
J Immunol. 2011 Mar 15;186(6):3472-83, PMID: 21321109, PMCID: 3127549.
|
|
Multiscale Granuloma Simulations
|
|
|
VI
|
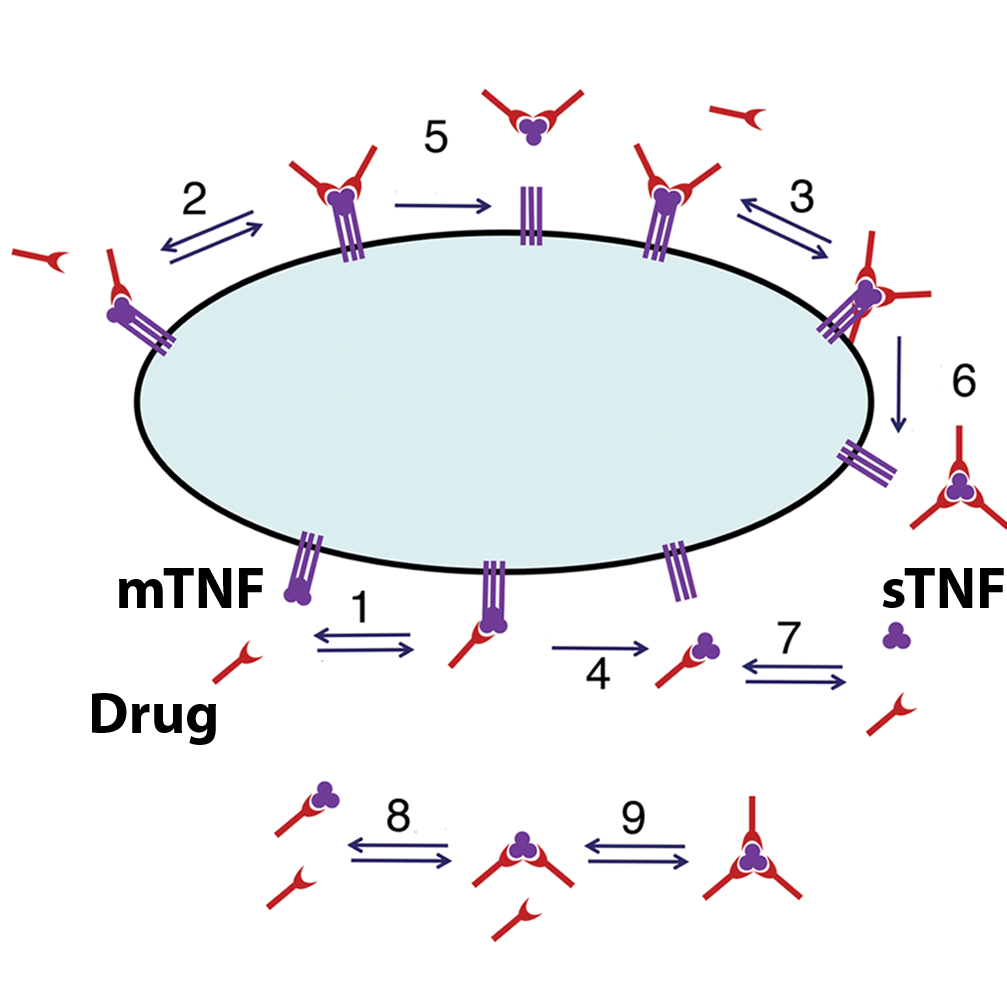
Differential risk of tuberculosis reactivation among anti-TNF
therapies is due to drug binding kinetics and permeability
|
|
|
Mohammad Fallahi-Sichani, JoAnne L. Flynn, Jennifer J. Linderman,
Denise E. Kirschner, The Journal of Immunology, 188, no. 7 (2012): 3169-78.,
PMID: 22379032, PMCID: 3311778
|
|
Multiscale Anti-TNF Drug Simulations
|
|
|
VII
|
Microenvironments in Tuberculous Granulomas Are
Delineated by Distinct Populations of Macrophage Subsets
and Expression of Nitric Oxide Synthase and Arginase
Isoforms
|
|
|
Joshua T. Mattila, Olabisi O. Ojo, Diane Kepka-Lenhart,
Simeone Marino, Denise E. Kirschner, Laura E. Via, Clifton E. Barry,
III, Korean MD, Philana Ling Lin, Edwin Klein, Sidney M. Morris
Jr., JoAnne L. Flynn,
J Immunol, July 15th 2013, 191, pages 773-784, PMID: 23749634, PMCID: 3746594
|
|
Multiscale M1-M2 Granuloma Simulations
|
|
|
VIII
|
Predicting lymph node output efficiency through systems biology
|
|
|
Chang Gong, Joshua T. Mattila, Mark Miller, JoAnne L. Flynn, Jennifer J. Linderman, D. Kirschner,
Journal of Theoretical Biology, Volume 335, October 21 2013, Pages 169-184, ePUB: June 29, 2013,
PMID: 23816876, PMCID: 3783027
|
|
3d LN Simulations
|
|
|
IX
|
A computational tool integrating host immunity with antibiotic dynamics to study tuberculosis treatment
|
|
|
Elsje Pienaar, Nicholas A. Cilfone, Philana Ling Lin, Veronique Dartois, Joshua T. Mattila, J. Russell Butler,
JoAnne L. Flynn, Denise E. Kirschner, Jennifer J. Linderman, Journal of Theoretical Biology (2015), pp. 166-179,
published online: 24-DEC-2014, DOI: 10.1016/j.jtbi.2014.11.021, PMID: 25497475, PMCID: 4332617
|
|
Antibiotic Treatment Simulations
|
|
|
X
|
A systems pharmacology approach towards the
design of inhaled formulations of rifampicin
and isoniazid for treatment of tuberculosis
|
|
|
Nicholas A. Cilfone, Denise E. Kirschner,
and Jennifer J. Linderman, 2015, Pharmacometrics and Systems Pharmacology, DOI: 10.1002/psp4.22, PMID: 26225241, PMCID: 4394619
|
|
Inhaled Antibiotic Treatment Simulations
|
|
|
XI
|
Deletion of TGF-β1 Increases Bacterial Clearance by Cytotoxic T Cells in a Tuberculosis Granuloma
|
|
|
Hayley C. Warsinske, Elsje Pienaar, Jennifer J. Linderman, Joshua T. Mattila, Denise E. Kirschner, 2017 (submitted), Frontiers in Immunology
|
|
TGF-β1 in a Tuberculosis Granuloma Simulations
|
|
|
XII
|
A computational model tracks whole-lung Mycobacterium tuberculosis infection and predicts factors that inhibit dissemination,
|
|
|
Timothy Wessler, Louis R. Joslyn, H. Jacob Borish, Hannah P. Gideon, JoAnne L. Flynn, Denise E. Kirschner, Jennifer J. Linderman, Online 24 July 2019, biorxiv.org, (accepted at PLoS Computational Biology)
|
|
Tuberculosis Granuloma Simulations
|
|
|
XIII
|
Neutrophil Dynamics Affect Mycobacterium tuberculosis Granuloma Outcomes and Dissemination,
|
|
|
Caitlin Hult, Joshua T. Mattila, Hannah P. Gideon, Jennifer J. Linderman, and Denise E. Kirschner, Front. Immunol., 05 October 2021
|
|
Neutrophil Simulations
|
|
|
XIV
|
A framework for multi-scale intervention modeling:
virtual cohorts, virtual clinical trials, and model-to-model comparisons,
|
|
|
Christian T. Michael, Sayed Ahmad Almohri, Jennifer J. Linderman, and Denise E. Kirschner,
Frontiers in Biology Systems, January 22, 2024, vol 3
|
|
Virtual Cohort Simulations
|
|